 |
Release version: v4.1 (September 15, 2025) |
|
Search PolyA_DB
|
||||
|
Description:
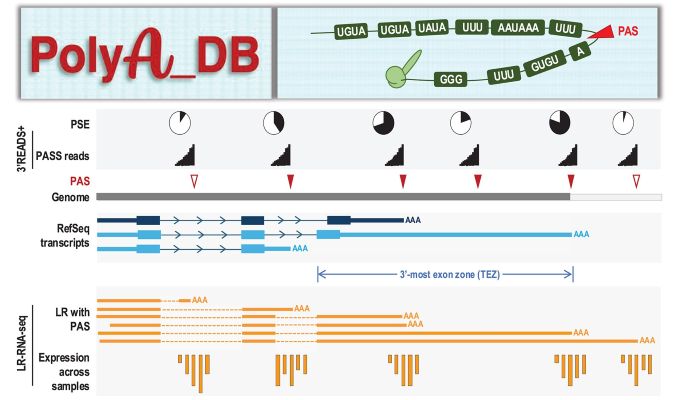
PolyA_DB version 4 catalogs polyA sites (PAS) in several mammalian genomes using both 3' end short-read RNA-seq (3'READS+) data and long-read RNA-seq (PacBio and Oxford Nanopore) data. Gene annotations are based on NCBI Gene and RefSeq databases. LR-RNA-seq data are used to connect PAS identified by 3'READS+ to genes and determine PAS types in the context of gene and splicing patterns. PolyA_DB v4 also contains PAS conservation and strength information. The data are linked to UCSC genome browser for visualization.
|
||||
|
References: 1. Yu S, Chen W, Wang L, Jewell S, Mammedova A, Han S, Wickramasinghe J, Barash Y, Tian B. (2025). PolyA_DB v4: systematic polyA site identification and isoform annotation in human and mouse genomes using 3’ end and long-read sequencing data. Nucleic Acids Res. 2025 Nov 29:gkaf1212. 2. Zheng D, Liu X, Tian B. (2016). 3'READS+, a sensitive and accurate method for 3' end sequencing of polyadenylated RNA. RNA.22:1631-9. 3. Hoque M*, Ji Z*, Zheng D, Luo W, Li W, You B, Park JY, Yehia G, Tian B. (2013). Analysis of alternative cleavage and polyadenylation by 3' region extraction and deep sequencing. Nature Methods 10:133-9. 4. Reese F et al. (2023) The ENCODE4 long-read RNA-seq collection reveals distinct classes of transcript structure diversity.bioRxiv 2023.05.15.540865. 5. Glinos DA et al. (2022) Transcriptome variation in human tissues revealed by long-read sequencing. Nature, 2022. 608:353-9 6. Mudge JM et al. (2025) GENCODE 2025: reference gene annotation for human and mouse. NAR, 2025. 53:D966-75 |
|
Please contact Shan Yu/Bin Tian for questions/comments/suggestions.
|